Abstract
BACKGROUND AND PURPOSE: High resolution 7T MRI is increasingly used to investigate hippocampal subfields in vivo, but most studies rely on manual segmentation which is labor intensive. We aimed to evaluate an automated technique to segment hippocampal subfields and the entorhinal cortex at 7T MRI.
MATERIALS AND METHODS: The cornu ammonis (CA)1, CA2, CA3, dentate gyrus, subiculum, and entorhinal cortex were manually segmented, covering most of the long axis of the hippocampus on 0.70-mm3 T2-weighted 7T images of 26 participants (59 ± 9 years, 46% men). The automated segmentation of hippocampal subfields approach was applied and evaluated by using leave-one-out cross-validation.
RESULTS: Comparison of automated segmentations with corresponding manual segmentations yielded a Dice similarity coefficient of >0.75 for CA1, the dentate gyrus, subiculum, and entorhinal cortex and >0.54 for CA2 and CA3. Intraclass correlation coefficients were >0.74 for CA1, the dentate gyrus, and subiculum; and >0.43 for CA2, CA3, and the entorhinal cortex. Restricting the comparison of the entorhinal cortex segmentation to a smaller range along the anteroposterior axis improved both intraclass correlation coefficients (left: 0.71; right: 0.82) and Dice similarity coefficients (left: 0.78; right: 0.77). The accuracy of the automated segmentation versus a manual rater was lower, though only slightly for most subfields, than the intrarater reliability of an expert manual rater, but it was similar to or slightly higher than the accuracy of an expert-versus-manual rater with ∼170 hours of training for almost all subfields.
CONCLUSIONS: This work demonstrates the feasibility of using a computational technique to automatically label hippocampal subfields and the entorhinal cortex at 7T MRI, with a high accuracy for most subfields that is competitive with the labor-intensive manual segmentation. The software and atlas are publicly available: http://www.nitrc.org/projects/ashs/.
ABBREVIATIONS:
- ASHS
- automated segmentation of hippocampal subfields
- CA
- cornu ammonis
- DSC
- Dice similarity coefficient
- DG
- dentate gyrus
- ERC
- entorhinal cortex
- ICC
- intraclass correlation coefficient
- SUB
- subiculum
The segmentation of subfields within the hippocampal formation on in vivo MRI is of major interest because these small anatomic subregions are potentially differentially affected in neuropsychiatric and neurologic disorders, including Alzheimer disease, major depressive disorder, posttraumatic stress disorder, and schizophrenia.1 In the previous decade, >20 segmentation protocols for MRI have been published for the hippocampal subfields and adjacent medial temporal lobe structures.2 Most of these protocols rely on manual segmentation,3⇓⇓⇓⇓⇓–9 which is labor-intensive, requires a long training period, and is often difficult to reproduce between research centers. Automated segmentation methods can help overcome these problems. To our knowledge, currently, only 4 automated segmentation methods exist,10⇓–12 3 of which were developed and evaluated on scans acquired at 3T MR imaging. Only the new FreeSurfer method (http://surfer.nmr.mgh.harvard.edu), developed by Iglesias et al,13 was developed by using a higher resolution 7T postmortem atlas set, though its application has only been demonstrated at lower field strengths. The advantage of in vivo 7T MRI is that high-resolution 3D images can be generated with a relatively short scanning time, making it possible to visualize hippocampal anatomy in greater detail.
Recently, an increasing number of 7T studies have been published on the hippocampal subregional morphology.14⇓–16 Several manual segmentation protocols exist for 7T MRI,5,7,17 and a semi automatic technique for measuring the thickness of hippocampal subfields and layers in the hippocampal body was developed by Kerchner et al.18 In this study, we evaluated the performance of a fully automated segmentation technique for labeling hippocampal subfields and the entorhinal cortex (ERC) at 7T MR imaging, which comes with a new set of challenges, including field inhomogeneity artifacts and increased image size. We do so by adapting a technique previously developed for 3T MRI12 to 7T MRI, labeled by using the manual annotation protocol developed by Wisse et al (2012).5 This protocol and the resulting automatic segmentation cover most of the longitudinal axis of the hippocampal formation. In addition, this article is the first to show that automatic segmentation performs competitively with interrater manual segmentation when the whole length of the hippocampus is labeled. Previously, only Yushkevich et al19 performed a comparison of automatic hippocampal subfield segmentation and interrater manual segmentation reliability, doing so at 3T and only in the body of the hippocampus.
Materials and Methods
Participants
Participants were included from the PREDICT-MR,16 an ancillary study to the PREDICT-NL study,20 which aimed to investigate determinants and consequences of brain changes on MR imaging in general practice attendees. The cohort included individuals 18 years of age or older who were asked to participate while in the waiting room of their general practitioner, irrespective of their symptoms.
The studies were performed in accordance with the principles of the Declaration of Helsinki and approved by the local ethics committee from the University Medical Center in Utrecht. Written informed consent was obtained from all participants.
Study Sample for the Atlas Set, Intrarater Reliability, and the Interrater Reliability Set
For the atlas set, 30 participants with a 7T T2-weighted MRI scan, required for the hippocampal subfield segmentation protocol, were randomly selected from the 47 participants in total. Images of 4 were considered to have relatively poor quality due to excessive subject motion, leaving 26 participants for the current study (mean age, 59 ± 9 years; 46% men; median Mini-Mental State Examination score,21 29; range, 25–30).
As a comparison for the reliability of the automated segmentation, we included overlap and reliability values of a single rater (L.E.M.W., rater 1; intrarater reliability) and of 2 raters (L.E.M.W., rater 1, and A.M.H., rater 2; interrater reliability). The intrarater reliability was established in a previous study,5 and the dataset consisted of the first 14 participants of the PREDICT-MR study (overlap with the atlas set, n = 7).5 For the interrater reliability, a random set of 14 MRI scans of PREDICT-MR was selected for segmentation (overlap with the atlas set, n = 12). The reliability analysis was after a training period of rater 2 of approximately 5 months, 1 day a week.
See On-line Fig 1 for a Venn diagram describing the samples.
Image Acquisition
All scans were performed on a 7T MR imaging scanner (Philips Healthcare, Best, the Netherlands) by using a volume transmit coil and a 16-channel receive coil (Nova Medical, Wilmington, Massachusetts) (participants included in the study later than May 2011 were scanned with a volume-transmit and 32-channel receive head coil [Nova Medical]). The 7T protocol included 0.70 × 0.70 × 0.70 mm3 3D T2-weighted TSE with a TR of 3158 milliseconds, a nominal TE of 301 milliseconds (with a contrast equivalent to a TE of 58 ms for brain tissue in spin-echo sequences with full refocusing angles), a flip angle of 120°(to partly compensate inhomogeneity in the radiofrequency field), a TSE factor of 182, a matrix size of 356 × 357 × 272, the application of 2D sensitivity encoding with acceleration factors of 2.0 × 2.8 (anterior-posterior × right-left), and a scan duration of 10 minutes and 15 seconds.5 The images were interpolated by zero-filling during reconstruction to a nominal spatial resolution of 0.35 × 0.35 × 0.35 mm3. Moreover, the 7T MRI protocol included a 1.00 × 1.00 × 1.00 mm3 T1-weighted sequence with a TR of 4.8 ms, TE of 2.2 ms, TI of 1240 ms, a TR of the inversion pulses of 3500 ms, a matrix size of 200 × 250 × 200, and a scan duration of 1 minute and 57 seconds.
Manual Segmentation
The cornu ammonis (CA) fields CA1, CA2, CA3 and the dentate gyrus (DG) (the dentate gyrus label includes both the granular cell layer of the dentate gyrus and the hilar region, sometimes called CA4), subiculum (SUB), and ERC were manually segmented, blinded to participant information, by using in-house-developed software22 based on MeVisLab (MeVis Medical Solutions, Bremen, Germany23). Segmentations were performed on coronal images, angulated perpendicular to the long axis of the hippocampal formation. The ERC was segmented according to the protocol by Goncharova et al,24 except for the posterior border, for which we followed the protocol of Insausti et al.25 CA1, CA2, CA3, DG, and SUB were segmented according to a previously published protocol,5 covering most of the long axis of the hippocampal formation. The anterior border was the most anterior section on which the hippocampus could be observed. The posterior border was defined as the section in which the total length of the fornix was visible. This was the most posterior section on which hippocampal subfields were segmented. Beyond this point, subfields fused together and could not be delineated reliably.
Automated Segmentation
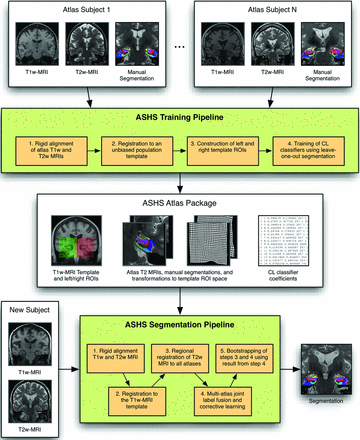
We applied the automated segmentation of hippocampal subfields (ASHS) technique by using this atlas set. Briefly, the method applies deformable registration of the T1- and T2-weighted images,26 multi-atlas joint label fusion,27 and voxelwise learning-based error correction,28 to propagate anatomic labels from a set of manually labeled training images to an unlabeled image. ASHS was evaluated by using a leave-one-out cross-validation (ie, when automatically segmenting the 7T scan of 1 participant in the study, the scans of the remaining 25 participants were used as training data). The resulting automatic segmentation was then compared with the manual segmentation of the same participant. Certain parameters of the method were modified for the 7T segmentation to account for differences in image size and resolution. More details are provided in Fig 1 and the On-line Appendix.
Training and segmentation pipelines in ASHS. Reprinted with permission from Yushkevich et al. 12 Copyright 2014 Wiley Periodicals.
Statistical Analyses
Volumes generated by manual and automated segmentations were compared by using a paired t test. The accuracy of automatic segmentation relative to manual segmentation (ASHS versus rater 1) was assessed in terms of relative overlap by using the Dice similarity coefficient (DSC).29 The DSC was computed separately for each subfield and jointly for all subfields (generalized DSC,30 see the On-line Appendix for a definition). The consistency of volume measurements derived from automatic and manual segmentations was measured by using the intraclass correlation coefficient (ICC) by using SPSS, Version 20 (IBM, Armonk, New York). The ICC variant that measured absolute agreement under a 2-way random analysis of variance model was used. Analogous statistical methods were used to compute the ICC and DSC between repeat segmentations of the same scans by rater 1 (intrarater reliability) and between 2 raters (rater 1 versus 2, interrater reliability).
In the 12 subjects who were included in the atlas set and the sample for the interrater reliability of the 2 manual raters, we performed additional analyses to test whether the DSCs of ASHS versus rater 1 were significantly different from the DSCs of rater 2 versus rater 1, by using Wilcoxon signed rank tests (2-sided).
In addition, we evaluated the ERC segmentation without the most anterior and posterior sections. We created a mask for the manual segmentation by removing the sections anterior to the head of the hippocampus and by removing the 4 most anterior and posterior sections of the resulting set of sections.
Results
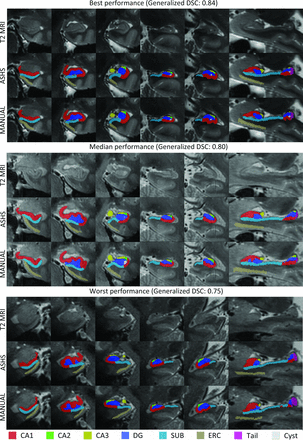
Figure 2 presents a visualization of the comparison of the automated and corresponding manual segmentation from the cross-validation experiment. Based on the generalized DSC, the best, median, and worst performances are shown. This figure shows that in the upper and middle panel (the best and median performance), the automated segmentations look very similar to the manual segmentations, though in the middle panel, small localized differences can be observed. For example, the segmentation of CA3 (yellow) and the ERC (light brown) is generally smaller/thinner in the automated-versus-manual segmentation. In the lower panel, showing the segmentation with the lowest generalized DSC, the overall location of the subfields is still similar in the manual and automated segmentation. However, local differences can be observed. For example, CA2 (green) and CA3 (yellow) are smaller in the automated-versus-manual segmentation. In addition, we observed that the mismatch occurs mainly in the segmentation of the most anterior sections for CA2, CA3, and the ERC. The automated segmentation of CA2, CA3, and the ERC included mostly fewer sections but sometimes more sections than the manual segmentation, which was likely a major source of inconsistency between the annotations. We will address this issue later in the “Results” for the ERC and in the “Discussion.” Figure 3 shows a 3D rendering of the automated segmentation of hippocampal subfields and the ERC.
Examples of results from the automated segmentation from the cross-validation experiment with the best (upper panel, left hemisphere), median (middle panel, left hemisphere), and worst performance (lower panel, right hemisphere). In each panel in the top row, the raw T2 image is shown; in the second row, the automated segmentation of hippocampal subfields is shown; and the third row, the manual segmentation is shown.
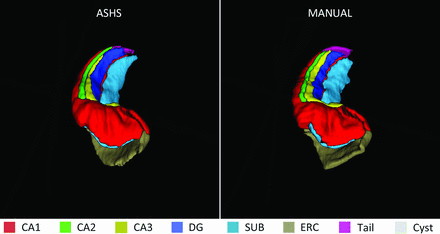
3D rendering of an automated (ASHS) and a manual segmentation.
Mean volumes of the manual and automated segmentation are shown in Table 1. CA1, DG, and SUB volumes generated by the automated segmentation were similar to those of manual segmentation, but CA2, CA3, and ERC volumes were smaller compared with the manual segmentation (P < .05). The DSC of ASHS versus rater 1 was >0.75 for the larger subfields CA1, DG, SUB, and ERC; however it was lower for the smaller subfields CA2 and CA3 (Table 2). The mean generalized DSC across all subfields in the left hemisphere was 0.80 ± 0.03, and for the right hemisphere, it was 0.79 ± 0.03. The ICC was >0.74 for the larger subfields CA1, DG, and SUB; however, it was lower for the ERC and the smaller subfields of CA2 and 3. Combining CA2 and 3 into a single label increased the bilateral DSC values and the right ICC compared with the segmentation of CA2 and CA3 alone.
Volumes of manual and automated segmentation
ICC and DSC among automated and corresponding manual segmentations, intrarater reliability of a single manual rater, and interrater reliability of 2 independent manual raters
Notably, the above results show a discrepancy between the ICC and the DSC values for the ERC. As described above, the automated segmentation of the ERC included mostly fewer sections, but sometimes more sections than the manual segmentation, which likely affected the ICC more than the DSC. We recalculated the ICC and DSC in a restricted range, as described in the “Materials and Methods” section, and found higher ICC values (left: 0.71, right: 0.82) and slightly higher DSC values (left: 0.78 ± 0.08; right: 0.77 ± 0.06).
Table 2 also shows the intrarater reliability of manual segmentation by rater 1.5 Overall, the intrarater reliability was higher than the agreement between the automated and manual segmentations. However, for automatic techniques such as ASHS that are trained on manual segmentations, the intrarater reliability of manual segmentation represents the theoretic upper bound for the agreement of automatic segmentation with manual segmentation. In addition, Table 2 shows the interrater reliability and overlap for 2 manual raters. The DSC values of ASHS versus rater 1 were higher for the larger subfields than the DSCs of rater 1 versus 2, and there were similar values for the smaller subfields. In additional analyses in the subjects who were included in both the atlas set and the set for the interrater reliability for the 2 manual raters, the DSC of ASHS versus rater 1 was significantly higher than the DSC of rater 1 versus 2 for the left ERC (P = .04), left and right SUB (P < .01; P < .01), right CA1 (P = .03), and left and right DG (P = .02; P < .01), and at a trend level for the right ERC (P = .08). It was equal for left CA1 (P = .14), left and right CA2 (P = .48; P = .58), and left CA3 (P = .43). Only for right CA3 was the DSC of the second rater higher at a trend level (P = .08) than that of ASHS. ASHS also had slightly higher or similar ICC values for most the subfields compared with the second rater, except for the DG, CA3, and right CA2.
Discussion
The current study demonstrates that automated segmentation of hippocampal subfields and the ERC at 7T MRI is feasible and that the errors of automatic segmentation are comparable with and in some cases even lower than the disagreement between 2 manual raters applying the same segmentation protocol. ASHS attained high accuracy (ICC > 0.74, DSC > 0.75) for larger subfields, including CA1, the DG, and SUB and lower accuracy for the ERC and smaller subfields, including CA2 and CA3. The anterior and posterior boundaries of the ERC were an important source of disagreement between the manual and automated segmentation. Restricting the range of ERC segmentation increased the accuracy, indicating that the ERC segmentation is accurate except at its anterior and posterior segments.
The high accuracy for the larger subfields, which is close to the intrarater reliability of this manual protocol,5 is promising and highly relevant, given the increasing number of sites using 7T MRI for hippocampal subfield research.5,14,17,31 The lower accuracy of the small subfields is consistent, to some extent, with that of the manual rater.5 It should be noted that small or thin structures are penalized by the DSC; as also mentioned by Pipitone et al,11 who showed that when comparing the automated segmentation with the manual segmentation shifted by 1 voxel, the DSCs of smaller structures were affected most.
As Table 1 shows, smaller structures (CA2, CA3, and ERC) were undersegmented by ASHS. The tendency of multiatlas label fusion algorithms to undersegment certain structures is a known limitation,32 and the machine learning corrective learning step in ASHS28 is meant to mitigate this effect, though it is not theoretically guaranteed to do so. In this study, corrective learning only partially reduced the undersegmentation error for CA2, CA3, and ERC (CA2 left: from 0.050 to 0.054; right: from 0.055 to 0.066; CA3 left: from 0.09 to 0.10; right: from 0.08 to 0.09; ERC left: from 0.46 to 0.47; right: from 0.47 to 0.49). As described in the “Results” section, the mismatch between the automated and manual method occurs mainly in the segmentation of the most anterior and posterior sections for CA2, CA3, and the ERC. This finding is not surprising, given that the anterior and posterior boundaries of CA2, CA3, and the ERC are based on a heuristic geometric rule rather than specific boundaries visible in the images. Restricting the range of the ERC indeed greatly increased the accuracy which is much closer to the intrarater reliability. In addition, the automated method slightly but systematically undersegments CA3 and the ERC in-plane. This undersegmentation might be a point for future improvement, for example, by incorporating a statistical shape or by manually retouching the automated segmentation of CA3. The reliability of the CA2 and CA3 segmentation warrants caution for future studies. Investigators might consider excluding these subfields from analyses or grouping them with either CA1 or the DG, depending on their research interests.
Notably, the automated segmentation performs similar or, in some cases, slightly better than a novice second rater for most of the subfields. Training a second rater takes considerable time in general, and specifically for this high-resolution data and detailed segmentation protocol, which includes several subfields and extends along most of the long axis of the hippocampus. The segmentation of one hippocampus can take up to 8 hours initially and 2 hours after 5 months of training. Training on the whole protocol can therefore take several months, underlining the need for an automated segmentation method. ASHS makes it feasible to perform automatic subfield segmentation and morphometry in large datasets, where manual segmentation by a single rater is prohibitive.
In the context of other automated segmentation methods,10⇓–12,33 the current method has a comparable and even slightly higher accuracy for the segmentation of almost all subfields. Only CA2 and 3 in the protocol of Van Leemput et al10 had higher accuracy values (DSC is approximately 0.09 higher). However, the segmentation protocol by Van Leemput et al has received considerable critiques,34,35 among others, on the placement of the boundaries that resulted in a larger CA2 and 3 volume in the Van Leemput protocol compared with our protocol. This probably explains the difference in DSC values. DSC values for the CA1, DG, and SUB were 0.03–0.28, 0.02–0.20, and 0.03–0.38 higher than those in prior studies,10⇓–12,33 most of which were performed at 3T MR imaging. For the smaller subfields CA2 and CA3 or the combined CA2+3, DSC values were 0.09–0.10, 0.01–0.05, and 0.23–0.25 higher than the DSC values of previous studies that used subfield boundaries comparable with those in the current study.11,12 Most interesting, the accuracy for segmenting hippocampal subfields in the current 7T study was slightly higher compared with a recent study using the same ASHS technique on anisotropic 3T data,12 despite the fact that the intrarater reliability of the 3T study was higher than that for the 7T study. This result indicates that there might be added value in using 7T data for the segmentation of hippocampal subfields.
The overlap and ICC values for the whole ERC are lower but approach the values of other automated segmentation methods.12,36,37 After restricting the range of the ERC segmentation, the accuracy improved and was well within the range of previous studies. This suggests that despite variability in the anterior and posterior boundary of the ERC, reliable measures of part of the ERC volume can be derived from ASHS segmentation. Another option for future work would be to manually correct the segmentation of the ERC, which would still take less time than a full segmentation.
A limitation of the current study, shared with all other published manual hippocampal subfield segmentation methods, is that in many cases, the actual anatomic boundaries between subfields cannot be inferred on in vivo MR imaging and are partly based on geometric rules. Resulting subfields may, therefore, include parts of neighboring regions. Another limitation is that ASHS is a computationally intensive method and requires >24 hours on a single central processing unit core to perform the segmentation of 1 participant. Furthermore, neither the current evaluation of ASHS nor the previous evaluation in Yushkevich et al12,19 has examined the ability of the ASHS atlases to generalize to scans obtained on different MR imaging scanners and with different MR imaging parameters. Considering that the MR imaging scanner and isotropic acquisition used in this study are used by very few research centers, it is unlikely that by directly using our atlas, other research groups will attain the same segmentation performance as reported in this article. However, ASHS is, by design, an adaptable technique and can be retrained by other groups by using different MR imaging protocols, provided that a set of manual segmentations is available. Moreover, in previous work, we have used atlases constructed by using MRI scans with one protocol to label medial temporal lobe subregions in scans obtained with a different protocol and field strength. For instance, we used an atlas developed on 4T MRI to investigate hippocampal subfields on 3T MRI and demonstrated stronger discrimination of CA1 compared with total hippocampal volume between those with prodromal Alzheimer disease and controls,38 but also showed that manual correction of ASHS results further improved discrimination of the CA1. Similarly, ASHS trained on data from a single 3T scanner was applied to multisite data from Alzheimer's Disease Neuroimaging Initiative 2 in Mueller et al,39 with sensible results. Although we have not validated the current 7T ASHS approach on other datasets, we have applied it on a few 0.4 × 0.4 × 1.0 mm3 7T scans obtained on a Siemens scanner (Siemens, Erlangen, Germany) with visually satisfactory segmentation results (see On-line Fig 2 for an example). In future work, it will be important to quantitatively evaluate the accuracy of ASHS in cross-scanner applications, as well as to measure how differences in the presence and severity of neurodegenerative disease in the atlas set and the target images affect segmentation accuracy. The fact that the current evaluation was performed in patients without known neurodegenerative disease is a limitation, though, in Yushkevich et al (2015),12 ASHS accuracy did not differ significantly between patients with mild cognitive impairment and controls. Finally, the datasets to evaluate the accuracy of ASHS versus rater 1 and the inter- and intrarater reliability of the manual raters only partially overlapped, which may have introduced a bias, though it should be noted that they were all drawn, without any consideration of image or segmentation quality, from the same study population and the scan quality in the resulting datasets was comparable among subjects. When comparing the DSCs of ASHS versus rater 1 with the DSCs for the intrarater reliability and the DSCs of ASHS versus rater 1 versus those of rater 1 versus 2 in the smaller, overlapping datasets, we saw no notable difference in the results (On-line Table). This finding indicates that the reliability of the segmentation was similar in all subjects and that the selection of scans probably did not introduce a bias.
Conclusions
We present a fully automated segmentation method of hippocampal subfields at 7T MRI with high accuracy for most of the subfields. The accuracy of this method is competitive with other published automated methods and with the interrater reliability for manual segmentation. Both the software and the atlas are publicly available at http://www.nitrc.org/projects/ashs/.
Acknowledgments
We acknowledge the use of MeVisLab by MeVis Medical Solutions, Bremen, Germany.
Footnotes
Paul A. Yushkevich and Mirjam I. Geerlings shared last authorship and contributed equally to this work.
Disclosures: David A. Wolk—UNRELATED: Consultancy: Piramal, Comments: consulting on the use of amyloid imaging in Alzheimer disease. Paul A. Yushkevich—RELATED: Grant: National Institutes of Health (AG037376)*; UNRELATED: Royalties: University of North Carolina, Chapel Hill,* Comments: I am on a patent for unrelated technology and receive royalties of about US $200.00 every year. Mirjam I. Geerlings—RELATED: Grant: Dutch Brain Foundation (Nederlandse Hersenstichting) project No. number 2012(1)-43.* *Money paid to the institution.
This work was funded by the National Institute on Aging, grant Nos. K23 AG028018, P30AG010124, and R01 AG037376; the National Institute of Biomedical Imaging and Bioengineering, grant Nos. R01 EB014346 and R01 EB017255. Hugo Kuijf was financially supported by the project Brainbox (quantitative analysis of MR brain images for cerebrovascular disease management), funded by the Netherlands Organisation for Health Research and Development in the framework of the research program Innovative Medical Devices Initiative, project 104002002. This research was also supported by a grant from the Dutch Brain Foundation (Hersenstichting Nederland: project No. 2012 [1]-43).
Indicates open access to non-subscribers at www.ajnr.org
References
- 1.
- 2.
- 3.
- 4.
- 5.
- 6.
- 7.
- 8.
- 9.
- 10.
- 11.
- 12.
- 13.
- 14.
- 15.
- 16.
- 17.
- 18.
- 19.
- 20.
- 21.
- 22.
- 23.
- 24.
- 25.
- 26.
- 27.
- 28.
- 29.
- 30.
- 31.
- 32.
- 33.
- 34.
- 35.
- 36.
- 37.
- 38.
- 39.
- Received July 31, 2015.
- Accepted after revision November 19, 2015.
- © 2016 by American Journal of Neuroradiology