Abstract
Objectives
The introduction of the 2016 WHO classification of CNS tumors has made the combined molecular and histopathological characterization of tumors a pivotal part of glioma patient management. Recent publications on radiogenomics-based prediction of the mutational status have demonstrated the predictive potential of imaging-based, non-invasive tissue characterization algorithms. Hence, the aim of this study was to assess the potential of multiparametric 18F-FET PET-MRI including MR fingerprinting accelerated with machine learning and radiomic algorithms to predict tumor grading and mutational status of patients with cerebral gliomas.
Materials and methods
42 patients with suspected primary brain tumor without prior surgical or systemic treatment or biopsy underwent an 18F-FET PET-MRI examination. To differentiate the mutational status and the WHO grade of the cerebral tumors, support vector machine and random forest were trained with the radiomics signature of the multiparametric PET-MRI data including MR fingerprinting. Surgical sampling served as a gold standard for histopathological reference and assessment of mutational status.
Results
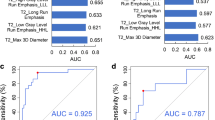
The 5-fold cross-validated area under the curve in predicting the ATRX mutation was 85.1%, MGMT mutation was 75.7%, IDH1 was 88.7%, and 1p19q was 97.8%. The area under the curve of differentiating low-grade glioma vs. high-grade glioma was 85.2%.
Conclusion
18F-FET PET-MRI and MR fingerprinting enable high-quality imaging-based tumor decoding and phenotyping for differentiation of low-grade vs. high-grade gliomas and for prediction of the mutational status of ATRX, IDH1, and 1p19q. These initial results underline the potential of 18F-FET PET-MRI to serve as an alternative to invasive tissue characterization.








Similar content being viewed by others
References
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, et al. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol (Berl). 2016;131:803–20.
Malone H, Yang J, Hershman DL, Wright JD, Bruce JN, Neugut AI. Complications following stereotactic needle biopsy of intracranial tumors. World Neurosurg. 2015;84:1084–9.
Pinker K, Shitano F, Sala E, Do RK, Young RJ, Wibmer AG, et al. Background, current role, and potential applications of radiogenomics. J Magn Reson Imaging JMRI. 2018;47:604–20.
Tian Q, Yan L-F, Zhang X, Zhang X, Hu Y-C, Han Y, et al. Radiomics strategy for glioma grading using texture features from multiparametric MRI. J Magn Reson Imaging JMRI. 2018;48;1518-28.
Akbari H, Bakas S, Pisapia JM, Nasrallah MP, Rozycki M, Martinez-Lage M, et al. In vivo evaluation of EGFRvIII mutation in primary glioblastoma patients via complex multiparametric MRI signature. Neuro-Oncol. 2018;20:1068–79.
Yu AC, Badve C, Ponsky LE, Pahwa S, Dastmalchian S, Rogers M, et al. Development of a combined MR fingerprinting and diffusion examination for prostate cancer. Radiology. 2017;283:729–38.
Gempt J, Soehngen E, Förster S, Ryang Y-M, Schlegel J, Zimmer C, et al. Multimodal imaging in cerebral gliomas and its neuropathological correlation. Eur J Radiol. 2014;83:829–34.
Kebir S, Weber M, Lazaridis L, Deuschl C, Schmidt T, Mönninghoff C, et al. Hybrid 11C-MET PET/MRI combined with “machine learning” in glioma diagnosis according to the revised glioma WHO Classification. Clin Nucl Med. 2018;44;214-20.
Singhal T, Narayanan TK, Jacobs MP, Bal C, Mantil JC. 11C-methionine PET for grading and prognostication in gliomas: a comparison study with 18F-FDG PET and contrast enhancement on MRI. J Nucl Med Off Publ Soc Nucl Med. 2012;53:1709–15.
Arita H, Kinoshita M, Kawaguchi A, Takahashi M, Narita Y, Terakawa Y, et al. Lesion location implemented magnetic resonance imaging radiomics for predicting IDH and TERT promoter mutations in grade II/III gliomas. Sci Rep. 2018;8:11773.
Leu K, Ott GA, Lai A, Nghiemphu PL, Pope WB, Yong WH, et al. Perfusion and diffusion MRI signatures in histologic and genetic subtypes of WHO grade II-III diffuse gliomas. J Neuro-Oncol. 2017;134:177–88.
Li Y, Liu X, Qian Z, Sun Z, Xu K, Wang K, et al. Genotype prediction of ATRX mutation in lower-grade gliomas using an MRI radiomics signature. Eur Radiol. 2018;28:2960–8.
Li Z-C, Bai H, Sun Q, Li Q, Liu L, Zou Y, et al. Multiregional radiomics features from multiparametric MRI for prediction of MGMT methylation status in glioblastoma multiforme: a multicentre study. Eur Radiol. 2018;28;3640-50.
Jiang Y, Ma D, Seiberlich N, Gulani V, Griswold MA. MR fingerprinting using fast imaging with steady state precession (FISP) with spiral readout. Magn Reson Med. 2015;74:1621–31.
Ma D, Coppo S, Chen Y, McGivney DF, Jiang Y, Pahwa S, et al. Slice profile and B1 corrections in 2D magnetic resonance fingerprinting. Magn Reson Med. 2017;78:1781–9.
Chung S, Kim D, Breton E, Axel L. Rapid B1+ mapping using a preconditioning RF pulse with TurboFLASH readout. Magn Reson Med. 2010;64:439–46.
Fedorov A, Beichel R, Kalpathy-Cramer J, Finet J, Fillion-Robin J-C, Pujol S, et al. 3D Slicer as an image computing platform for the quantitative imaging network. Magn Reson Imaging. 2012;30:1323–41.
Haury A-C, Gestraud P, Vert J-P. The influence of feature selection methods on accuracy, stability and interpretability of molecular signatures. PLoS ONE [Internet]. 2011;6. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3244389/.
Brown G, Pocock A, Zhao M-J, Luján M. Conditional likelihood maximisation: a unifying framework for information theoretic feature selection. J Mach Learn Res. 2012;13:27–66.
Caruana R, Karampatziakis N, Yessenalina A. An empirical evaluation of supervised learning in high dimensions. Proc 25th Int Conf Mach Learn ICML. 2008;1;96-103.
Aerts HJWL, Velazquez ER, Leijenaar RTH, Parmar C, Grossmann P, Carvalho S, et al. Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun. 2014;5:4006.
Shofty B, Artzi M, Ben Bashat D, Liberman G, Haim O, Kashanian A, et al. MRI radiomics analysis of molecular alterations in low-grade gliomas. Int J Comput Assist Radiol Surg. 2018;13:563–71.
Badve C, Yu A, Dastmalchian S, Rogers M, Ma D, Jiang Y, et al. MR fingerprinting of adult brain tumors: initial experience. AJNR Am J Neuroradiol. 2017;38:492–9.
Ma D, Gulani V, Seiberlich N, Liu K, Sunshine JL, Duerk JL, et al. Magnetic resonance fingerprinting. Nature. 2013;495:187–92.
Mehta BB, Coppo S, McGivney DF, Hamilton JI, Chen Y, Jiang Y, et al. Magnetic resonance fingerprinting: a technical review. Magn Reson Med. 2019;81:25–46.
Panda A, Mehta BB, Coppo S, Jiang Y, Ma D, Seiberlich N, et al. Magnetic resonance fingerprinting-an overview. Curr Opin Biomed Eng. 2017;3:56–66.
Suchorska B, Unterrainer M, Biczok A, Sosnova M, Forbrig R, Bartenstein P, et al. 18F-FET-PET as a biomarker for therapy response in non-contrast enhancing glioma following chemotherapy. J Neuro-Oncol. 2018;139:721–30.
Kebir S, Weber M, Lazaridis L, Deuschl C, Schmidt T, Mönninghoff C, et al. Hybrid 11C-MET PET/MRI combined with “machine learning” in glioma diagnosis according to the revised glioma WHO classification 2016. Clin Nucl Med. 2019;44:214–20.
Verger A, Stoffels G, Bauer EK, Lohmann P, Blau T, Fink GR, et al. Static and dynamic 18F-FET PET for the charactserization of gliomas defined by IDH and 1p/19q status. Eur J Nucl Med Mol Imaging. 2018;45:443–51.
Verger A, Filss CP, Lohmann P, Stoffels G, Sabel M, Wittsack HJ, et al. Comparison of 18F-FET PET and perfusion-weighted MRI for glioma grading: a hybrid PET/MR study. Eur J Nucl Med Mol Imaging. 2017;44:2257–65.
You S-H, Choi SH, Kim TM, Park C-K, Park S-H, Won J-K, et al. Differentiation of high-grade from low-grade astrocytoma: improvement in diagnostic accuracy and reliability of pharmacokinetic parameters from DCE MR imaging by using arterial input functions obtained from DSC MR imaging. Radiology. 2017;286:981–91.
Hsu CC-T, Watkins TW, Kwan GNC, Haacke EM. Susceptibility-weighted imaging of glioma: update on current imaging status and future directions. J Neuroimaging Off J Am Soc Neuroimaging. 2016;26:383–90.
Verger A, Taieb D, Guedj E. Is the information provided by amino acid PET radiopharmaceuticals clinically equivalent in gliomas? Eur J Nucl Med Mol Imaging. 2017;44:1408–10.
Verger A, Metellus P, Sala Q, Colin C, Bialecki E, Taieb D, et al. IDH mutation is paradoxically associated with higher 18F-FDOPA PET uptake in diffuse grade II and grade III gliomas. Eur J Nucl Med Mol Imaging. 2017;44:1306–11.
Bette S, Gempt J, Delbridge C, Kirschke JS, Schlegel J, Foerster S, et al. Prognostic value of O-(2-[18F]-fluoroethyl)-L-tyrosine-positron emission tomography imaging for histopathologic characteristics and progression-free survival in patients with low-grade glioma. World Neurosurg. 2016;89:230–9.
Lopci E, Riva M, Olivari L, Raneri F, Soffietti R, Piccardo A, et al. Prognostic value of molecular and imaging biomarkers in patients with supratentorial glioma. Eur J Nucl Med Mol Imaging. 2017;44:1155–64.
Zhang B, Chang K, Ramkissoon S, Tanguturi S, Bi WL, Reardon DA, et al. Multimodal MRI features predict isocitrate dehydrogenase genotype in high-grade gliomas. Neuro-Oncol. 2017;19:109–17.
Arbizu J, Tejada S, Marti-Climent JM, Diez-Valle R, Prieto E, Quincoces G, et al. Quantitative volumetric analysis of gliomas with sequential MRI and 11C-methionine PET assessment: patterns of integration in therapy planning. Eur J Nucl Med Mol Imaging. 2012;39:771–81.
Acknowledgments
We would like to thank the MR fingerprinting team Dres, M. Nittka, J. Pfeuffer, and G. Koerzdoerfer, Siemens Healthcare, Erlangen, for providing the prototype MRF sequence.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: Johannes Haubold, Lale Umutlu, Aydin Demircioglu. Performed the experiments: Johannes Haubold, Aydin Demircioglu, Lale Umutlu. Analyzed the data: Aydin Demircioglu, Johannes Haubold, Lale Umutlu, Marcel Gratz, Felix Nensa. Contributed reagents/materials/analysis tools: Martin Glas, Karsten Wrede, Ulrich Sure, Gerald Antoch, Kathy Keyvani, Mathias Nittka, Stephan Kannengiesser, Vikas Gulani, Mark Griswold, Ken Herrmann, Michael Forsting, Felix Nensa. Wrote the paper: Johannes Haubold, Lale Umutlu, Aydin Demircioglu. Patient recruitments: Karsten Wrede, Martin Glas. Manuscript editing: Johannes Haubold, Lale Umutlu, Aydin Demircioglu, Marcel Gratz, Martin Glas, Karsten Wrede, Ulrich Sure, Gerald Antoch, Kathy Keyvani, Mathias Nittka, Stephan Kannengiesser, Vikas Gulani, Mark Griswold, Ken Herrmann, Michael Forsting, Felix Nensa. Statistics: Aydin Demircioglu. The manuscript has been seen and approved by all authors.
Corresponding author
Ethics declarations
This study was conducted in accordance with all guidelines set forth by the approving institutional review board of the University Hospital Essen. All examinations were performed after written informed consent was obtained from all patients.
Conflict of interest
There was no conflict of interest apart from the in the disclaimer mentioned funding of the study.
Disclaimer
This study was partially supported by Siemens Healthcare. Siemens Healthcare had no role in the study design, data collection, data analysis, data interpretation, or writing of the report.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Oncology – Brain.
Electronic supplementary material
ESM 1
(DOCX 27 kb)
Rights and permissions
About this article
Cite this article
Haubold, J., Demircioglu, A., Gratz, M. et al. Non-invasive tumor decoding and phenotyping of cerebral gliomas utilizing multiparametric 18F-FET PET-MRI and MR Fingerprinting. Eur J Nucl Med Mol Imaging 47, 1435–1445 (2020). https://doi.org/10.1007/s00259-019-04602-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-019-04602-2