Abstract
BACKGROUND: Brain imaging in diffuse glioma is used for diagnosis, treatment planning, and follow-up.
PURPOSE: In this meta-analysis, we address the diagnostic accuracy of imaging to delineate diffuse glioma.
DATA SOURCES: We systematically searched studies of adults with diffuse gliomas and correlation of imaging with histopathology.
STUDY SELECTION: Study inclusion was based on quality criteria. Individual patient data were used, if available.
DATA ANALYSIS: A hierarchic summary receiver operating characteristic method was applied. Low- and high-grade gliomas were analyzed in subgroups.
DATA SYNTHESIS: Sixty-one studies described 3532 samples in 1309 patients. The mean Standard for Reporting of Diagnostic Accuracy score (13/25) indicated suboptimal reporting quality. For diffuse gliomas as a whole, the diagnostic accuracy was best with T2-weighted imaging, measured as area under the curve, false-positive rate, true-positive rate, and diagnostic odds ratio of 95.6%, 3.3%, 82%, and 152. For low-grade gliomas, the diagnostic accuracy of T2-weighted imaging as a reference was 89.0%, 0.4%, 44.7%, and 205; and for high-grade gliomas, with T1-weighted gadolinium-enhanced MR imaging as a reference, it was 80.7%, 16.8%, 73.3%, and 14.8. In high-grade gliomas, MR spectroscopy (85.7%, 35.0%, 85.7%, and 12.4) and 11C methionine–PET (85.1%, 38.7%, 93.7%, and 26.6) performed better than the reference imaging.
LIMITATIONS: True-negative samples were underrepresented in these data, so false-positive rates are probably less reliable than true-positive rates. Multimodality imaging data were unavailable.
CONCLUSIONS: The diagnostic accuracy of commonly used imaging is better for delineation of low-grade gliomas than high-grade gliomas on the basis of limited evidence. Improvement is indicated from advanced techniques, such as MR spectroscopy and PET.
ABBREVIATIONS:
- hsROC
- hierarchic summary receiver operating characteristic
- ROC
- receiver operating characteristic
- STARD
- Standard for Reporting of Diagnostic Accuracy
- CI
- credibility interval
- TP
- true-positive
- TN
- true-negative
- FP
- false-positive
- FN
- false-negative
Diffuse gliomas are the most common primary brain tumors in adults, with an annual incidence of approximately 6 per 100,000. Despite advances in neurosurgery, radiation therapy, and chemotherapy, gliomas are fatal.1 Brain imaging is indispensable for diagnosis, treatment planning, evaluation, and follow-up. Although imaging standards to plan resection and radiation therapy vary between institutions and specialists, conventional imaging is in common use, typically consisting of T1-weighted MR imaging before and after gadolinium and T2/FLAIR-weighted imaging for gliomas. Of these conventional sequences, T2/FLAIR-weighted imaging is often considered as a reference for low-grade gliomas, and T1-weighted gadolinium-enhanced imaging, for high-grade gliomas in neurosurgical planning, combined with T2-weighted imaging in radiation therapy planning.2,3
Compared with other cancer types, accurate delineation of gliomas within the brain for treatment planning is particularly important due to the proximity of eloquent brain structures, which are vulnerable to surgery and radiation therapy.4 Conversely, more extensive resections and boosted radiation therapy correlate with longer survival.5⇓–7 At the same time, clinical observations challenge the diagnostic accuracy of current imaging protocols: Gliomas recur even after a radiologically complete resection,8,9 and glioma cells have been detected outside MR imaging abnormalities.10,11 Brain imaging techniques, such as multivoxel spectroscopy and PET, were developed to improve tumor grading and delineation.12,13
Inherent in any regional treatment, such as surgery and radiation therapy, is the need to delineate a target volume, which mandates a dichotomous classification into tumor and normal tissue. Low- and high-grade gliomas have different treatment strategies and prognosis, while both are characterized by diffuse tumor infiltration. This supports our pooled analysis for diffuse glioma in addition to subgroup analysis by glioma grade. More accurate glioma delineation may improve the consistency between treatment results and survival. For instance, more accurate delineation may serve to identify patients eligible for more aggressive surgery than would have been considered on the basis of conventional imaging and may identify patients with glioma infiltration beyond meaningful surgical therapy so that useless and possibly harmful resections can be avoided.
The diagnostic accuracy of imaging techniques to delineate gliomas has not been systematically addressed, to our knowledge. In this meta-analysis, we estimate and compare the diagnostic accuracies of conventional imaging techniques and advanced MR imaging and PET to delineate newly diagnosed diffuse gliomas within brain tissue in adults.
Materials and Methods
Search Strategy
We aimed to identify all publications reporting glioma imaging correlated with histopathology for the sampled locations. Our data sources were the National Library of Medicine (PubMed/MEDLINE, beginning in 1966) and the Excerpta Medica DataBASE (EMBASE, beginning in 1947), accessed on February 29, 2016, searching MeSH and Emtree subject headings (On-line Appendix: Methods 1). The publication language was restricted to Western languages; the publication date was restricted until January 1, 2016. References of identified studies were reviewed for further eligible publications in adherence with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement guidelines (http://prisma-statement.org/).
Study Selection Criteria
Studies were eligible for further analysis on the basis of 5 inclusion criteria: First, an adult population or subpopulation was required with patients at least 18 years of age. Second, only newly diagnosed diffuse gliomas of World Health Organization grades II–IV were included to avoid imaging artifacts from previous treatment. Third, brain imaging of any technique was allowed as a diagnostic test. Fourth, histopathology of tissue samples was required as a reference test. Fifth, histopathology of samples and imaging test measurements had to be directly correlated by surgical navigation with 3D coordinates. When duplicate reports on the same population were retrieved, only the report with the most complete data was used for analysis. Studies also including patients of pediatric age or pathology other than diffuse glioma were included if the population of interest could be extracted from the data. Postmortem studies were excluded to avoid bias from end-stage disease. For interobserver agreement on study inclusion, a κ statistic was used.14
Quality and Outcome Measures
Any study reporting glioma imaging correlated with histopathology was independently assessed in full text for quality criteria by 2 observers (N.V. and F.W.A.H, with 3 and 8 years of experience in clinical neurosurgery) using the Standard for Reporting of Diagnostic Accuracy (STARD) guidelines, a 25-item checklist to explore reporting quality.15 Disagreements were resolved through adjudication by a third observer (P.C.D.W.H., with 15 years of experience in clinical neurosurgery). For interobserver agreement on quality assessment, the intraclass correlation coefficient was calculated. One observer extracted the study data (N.V.) on the imaging techniques, the histopathologic examination method, the colocalization method between imaging and histopathology, and the number of patients and samples categorized by glioma grades. The extracted data were verified by another observer (P.C.D.W.H.). Biopsy sample locations were categorized for each imaging technique as normal or abnormal signal according to the authors' test positivity criteria and as glioma or normal brain in correlated histopathologic examination according to the authors' definitions. We aimed to include individual sample data as much as possible, either by availability from the publication, by request for original data to corresponding authors (up to 3 times in case of nonresponse), or by estimation of data points from plots of image measurements versus tumor characteristics. If individual sample data were unavailable, we included the aggregated data as summaries of true-positive (TP), true-negative (TN), false-positive (FP), and false-negative (FN) samples. Accordingly, each imaging technique in every study provided at least 1 estimate of diagnostic accuracy.
Statistical Analysis
We used hierarchic summary receiver operating characteristic (hsROC) curve analysis with random effects for within-study and between-study variation to summarize estimates of the diagnostic accuracy for each imaging technique with >1 study available (On-line Appendix: Methods 2).16 The available data allowed hierarchic analysis of samples within techniques. The estimates with 95% credibility intervals (CIs) consisted of the summary true-positive rate (sensitivity: TP / [TP + FN]), the summary false-positive rate (1-specificity: FP/ [FP + TN]), the diagnostic odds ratio ([TP/FN]/[FP/TN]),17 and the summary area under the receiver operating characteristic (ROC) curve. Differences in estimates between imaging techniques in relation to reference imaging were also calculated and considered significant when the CI excluded zero. A Bayesian Markov chain Monte Carlo algorithm with 3 chains modeled the hsROC curve with the CI for each imaging technique on the basis of 5 parameters: the accuracy and its precision, the cut-point and its precision, and the scale parameter.18 The accuracy and the cut-point were allowed to vary among studies; the scale was allowed to vary among imaging techniques. These parameters provide an hsROC curve with an operating point on the curve representing the summary true-positive and false-positive rates. A Bayesian hsROC analysis was used for the following reasons: First, this approach is considered the standard for meta-analysis of diagnostic accuracy, taking into account the correlation between the true-positive and false-positive rates.18,19 Second, individual sample data and aggregated data, if individual data are unavailable, can be analyzed together per imaging technique to estimate its diagnostic accuracy.20,21 Third, varying test positivity criteria for imaging in studies can be accommodated.22 Fourth, partly missing data can be handled with this model. For the meta-analysis, vague priors were chosen for the prior distributions, so that the results primarily reflect inference from the presented data without prior knowledge. Normal distributions with mean of zero and variance of 10,000 represented the accuracy, cut-point and scale parameters, and inverse γ distributions, with a shape and rate of 0.01 representing the precisions. Uniform distributions were used as alternative vague priors. Summary estimates consisted of the median values with CIs of the posterior distributions. For the analysis, we used JAGS software, Version 4.0.1 (Jags Software; Newark, Delaware; http://mcmc-jags.sourceforge.net) from the rjags package (Version 4–3) for R statistical and computing software (http://www.r-project.org/). Sampling traces and distributions and Gelman-Rubin diagnostics were evaluated for evidence against convergence by using the coda package (Version 0.17.1; https://www.rdocumentation.org/packages/coda/versions/0.17-1).
Statistical tests for heterogeneity between studies are unavailable for diagnostic accuracy meta-analysis; therefore, heterogeneity was explored with subgroup analysis.23 Sensitivity analysis included analysis of the following: 1) the subset of higher quality studies, considered those with a prospective design, including quantified sample data in at least 15 patients and qualifying for the methodologic description of both neuroimaging and histopathology24; 2) the subset of studies with individual patient data; and 3) alternative vague prior distributions. Subgroups of low-grade (World Health Organization grade II) and high-grade (World Health Organization grade III or IV) gliomas were reported separately. Publication bias was explored graphically.
Results
The search strategy identified 8558 unique citations (Fig 1), of which 272 full-text publications were assessed for eligibility. Subsequently, 61 articles were included for meta-analysis on the basis of the 5 selection criteria. The interobserver agreement for inclusion was moderate (κ = 0.47; 95% CI, 0.37–0.57). A total of 3532 samples with correlated histopathologic examinations and imaging were included from 1309 patients with gliomas. For subgroup analysis by glioma grade, data could be extracted for 907 samples in 421 patients with low-grade gliomas and for 1380 samples in 814 patients with high-grade gliomas. Glioma subtypes, such as astrocytoma or oligodendroglioma, could not be analyzed in subgroups because data for these subgroups could usually not be extracted. Individual sample data were available from 19 studies. The higher quality subset consisted of 29 studies.
Flow chart of publication selection. The asterisk indicates that multiple exclusion criteria could apply to studies.
The mean STARD score of the included studies was 13 ± 2.8. The interobserver agreement for STARD quality assessment was substantial (intraclass coefficient = 0.66; 95% CI, 0.49–0.78).
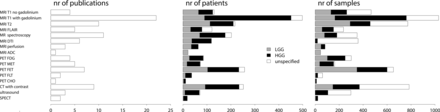
The included studies reported on 16 imaging techniques, including T1-weighted imaging before and after gadolinium, T2-, T2/FLAIR-, perfusion- and diffusion-weighted imaging (apparent diffusion coefficient), MR spectroscopy (choline to N-acetylaspartate ratio), diffusion tensor imaging (fractional anisotropy), and PET with these tracers: FDG, 11C methionine (MET), 18F fluoroethyltyrosine (FET), 18F fluorthymidine, or 18F or 11C choline (Cho). Imaging protocols varied widely; for instance, 22 studies used 1.5T MR imaging field strength, 15 used 3T, 1 used 0.15T; and field strength was unspecified in 11 studies. The number of studies, patients, and samples for each imaging technique categorized by glioma grades is plotted in Fig 2.
Number of publications, patients, and samples for imaging techniques categorized by glioma grades. LGG indicates low-grade glioma; HGG, high-grade glioma; FLT, 18F fluorthymidine; nr, number.
The reference standard to distinguish tumor and normal brain in tissue samples for all studies was microscopic examination with hematoxylin-eosin staining and immunohistochemical analysis. Nine studies reported the reference standard as the labeling index of proliferating cells; 6 studies reported the reference standard as the tumor infiltration index; and 2 studies, as the cellularity index. We followed the authors' definition to differentiate normal brain and glioma.
The method to correlate the histopathology with imaging was frameless stereotactic needle biopsies in 27 studies, frame-based stereotactic needle biopsies in 14, neuronavigated resection biopsies in 12, and unspecified stereotactic needle biopsies in 8.
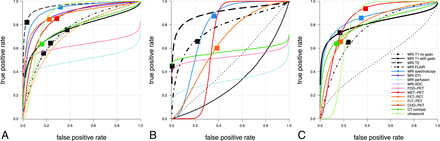
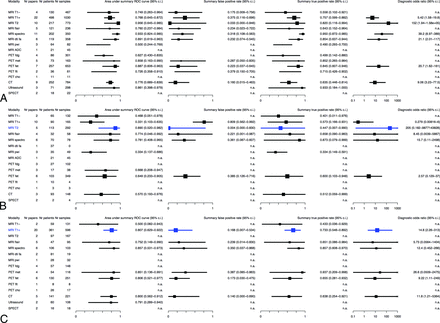
The hsROC curves of imaging techniques for diffuse glioma as a whole are plotted in Fig 3A, and for the subgroups of low- and high-grade gliomas, in Fig 3B, -C, respectively. The characteristics of these hsROC curves are listed in Fig 4.
Summary hierarchic ROC curves of imaging techniques. Estimated summary ROC curves are plotted by imaging technique in ROC space for diffuse gliomas as a whole—that is, low- and high-grade gliomas combined (A), low-grade glioma (B), and high-grade glioma (C). The operating points of curves represent the estimated summary true-positive and false-positive rates as indicated by solid squares. Solid squares are missing when convergence is not reached. Color coding shows conventional MR imaging in black, advanced MR imaging in shades of blue, PET imaging in shades of red, and CT and sonography imaging in shades of green.
Diagnostic accuracy characteristics of imaging techniques. Diagnostic performance of imaging techniques expressed as area under the curve, summary false-positive rate, summary true-positive rate, and diagnostic odds ratio for diffuse gliomas as a whole—that is, low- and high-grade gliomas combined (A), low-grade glioma (B), and high-grade glioma (C). The medians of posterior distributions are plotted as squares proportional to the number of observations with 95% credibility intervals. The most common clinically used imaging technique is in blue as reference. n.a. indicates nonavailable data because either <3 studies were available or the posterior samples did not converge; fa, fractional anisotropy.
The estimates of false-positive rates need to be interpreted with caution due to relative lack of data on true-negative samples. This “underrepresentation” of true-negatives and consequently bias in false-positive rates may be unbalanced across imaging techniques, creating additional biases when comparing the ROC curves.
In all gliomas considered, the area under the curve was highest for T2-weighted imaging (95.6%), followed by MR spectroscopy (93.3%). The false-positive rate was lowest in T2-weighted imaging (3.3%), followed by CT (16.0%); and the true-positive rate was highest for MR spectroscopy (95.0%), followed by sonography (93.3%). The diagnostic odds ratio was highest for T2-weighted imaging (152), followed by MR imaging spectroscopy (39.2).
For low-grade gliomas, the area under the curve was highest for T2-weighted imaging (89.0%), followed by MR spectroscopy (78.1%) and T2/FLAIR-weighted imaging (77.4%). The false-positive rate was lowest in T2-weighted imaging (0.4%), followed by T2/FLAIR-weighted imaging (22.1%); and the true-positive rate was highest for MR spectroscopy (87.3%), followed by T2/FLAIR-weighted imaging (65.8%). The diagnostic odds ratio was highest for T2-weighted imaging (205), followed by MR spectroscopy (15.7). Not surprising, the area under the curve was statistically significantly lower for T1-weighted gadolinium-enhanced imaging compared with T2-weighted imaging (difference of 54.4%; 95% CI, 8.4%–80.8%) and for perfusion-weighted imaging (difference of 54.1%; 95% CI, 7.8%–78.2%). Of note, T2-weighted imaging had a higher area under the curve, false-positive ratio, and diagnostic odds ratio in comparison with T2/FLAIR-weighted imaging, the other common clinically used MR imaging sequence.
For high-grade gliomas, the area under the curve was highest for MR spectroscopy (85.7%), followed by MET-PET (85.1%). The false-positive rate was lowest in CT (14.0%), followed by T1-weighted gadolinium-enhanced imaging (16.8%); and the true-positive rate was highest for MET-PET (93.7%), followed by MR spectroscopy (85.7%). The diagnostic odds ratio was highest for MET-PET (26.6), followed by T1-weighted gadolinium-enhanced imaging (14.8). MET-PET and MR spectroscopy had a higher area under the curve and true-positive rate in comparison with the common clinically used T1-weighted gadolinium-enhanced MR imaging (respectively, 80.7% and 73.3%). This came at the expense of higher false-positive rates for MET-PET and MR spectroscopy, respectively 38.7% and 35.0%. Most remarkable, CT with contrast had an area under the curve (80.0%) like that of T1-weighted gadolinium-enhanced MR imaging.
Sensitivity analysis demonstrated robustness against publication quality, individual sample data, and alternative vague priors (On-line Fig 1). Because statistical tests for heterogeneity were unavailable to take the association between sensitivity and specificity into account,23 we inspected the plots of the study data and hsROC curve for each technique (On-line Fig 2), demonstrating considerable heterogeneity among studies, resulting in a relatively large CI. For many imaging techniques, the number of publications was too small to exclude publication bias (On-line Fig 3). Small studies with small areas under the curve or low false-positive rates may be missing for T1-weighted gadolinium-enhanced imaging and T2-weighted imaging.
Discussion
The main findings of this meta-analysis for the diagnostic accuracy of neuroimaging to delineate diffuse gliomas are the following: 1) It is best with T2-weighted imaging, followed by MR spectroscopy. 2) It is better for low-grade gliomas with T2-weighted imaging than for high-grade gliomas with T1-weighted gadolinium-enhanced MR imaging; considering the area under the curve (89.0% versus 80.7%) and the diagnostic odds ratio (205 versus 14.8). 3) it may be improved in high-grade gliomas with MR spectroscopy or MET-PET. 4) It is not superior with T2/FLAIR-weighted imaging for low-grade gliomas. 5) It is not inferior with CT with contrast for high-grade gliomas. 6) It varies considerably between imaging techniques and shows heterogeneity between studies.
Thresholds for acceptable diagnostic accuracy of tumor imaging are undetermined. The accuracy of imaging for glioma delineation by MR imaging is, for instance, less than that for lesion detection in hepatocellular carcinoma by sonography, CT, or MR imaging.25 It is comparable with detection of metastatic lymph nodes in non-small cell lung cancer by CT or MR imaging, but it is less than that of PET.26,27 Moreover, it is less than that of the diagnosis of breast cancer by MR imaging.28 These studies, however, address the radiologic diagnosis of cancer by imaging, not the delineation of infiltrative cancer within normal tissue. The identification of cancer cells within normal-appearing imaging regions seems to be specific for glioma.10,29 We did not find meta-analyses of tumor delineation in other solid cancers.
The variation in diagnostic performance may be explained by the notoriously difficult diagnostic problem of delineating glioma cells, which gradually infiltrate brain tissue. Therefore, the concept of delineating a tumor by the presence or absence of cancer cells, on which ROC analysis is based, may oversimplify gradual glioma infiltration. Nevertheless, treatment target volumes are required for patient care.
Several factors may contribute to the observed variation in diagnostic performance. First, the scan protocols have not been standardized for any of these sequences. For the diagnostic standard of MR imaging, for instance, variation exists in scanner equipment, quality assessment and control, acquisition protocols, image processing, quantification, and interpretation by radiologists. Second, histopathologic examinations may vary due to incomplete sampling of heterogeneous tumors and interpretation differences among neuropathologists.30⇓–32 Third, the correlation between imaging measurements and histopathologic examination may be another source of variation. This colocalization depends on the precision of the navigated locations of tissue samples. Navigation precision has been found to be within several millimeters,33,34 whereas tissue may be heterogeneous at smaller distances.35
Improvement of diagnostic accuracy to delineate gliomas for regional therapy requires an offset between increasing the true-positive rate and decreasing the false-positive rate. Increasing the true-positive rate may be preferable for tumor control, whereas decreasing the false-positive rate may be preferable for preservation of functional integrity. From the perspective of tumor control, the overestimation of diffuse glioma (ie, the inadvertent declaration of normal brain as diffuse glioma) would be more acceptable than underestimation. Nevertheless, this is only acceptable when at the same time, surgery aims to minimize neurologic deficits from removal of critical brain regions whether or not they are infiltrated by tumor. This is usually done by brain mapping of functions with the patient under local anesthesia to push the resection to the functional limits.4 In other words, a more sensitive imaging delineation would be even more likely to require functional brain mapping as a safeguard against removal of critical brain regions potentially infiltrated by tumor. In this perspective, MR spectroscopy and PET might have the potential to increase the true-positive rate of glioma delineation for surgical strategies. However, a more sensitive tumor delineation should probably not prompt larger high-dose radiation fields because a similar safeguard against cognitive decline from radiation therapy is unavailable. Perhaps high-dose radiation therapy should rather focus on regions at high risk for tumor progression, whereas a lower dose could be acceptable in regions with a low risk for tumor progression.
Our observations may challenge current care standards. First, T2/FLAIR-weighted imaging has been proposed as the standard for radiologic response measurements in low-grade gliomas,3 whereas our data indicate that T2-weighted imaging has better diagnostic performance before treatment than T2/FLAIR-weighted imaging. Second, MR imaging, including T1-weighted gadolinium-enhanced and T2-weighted imaging, is considered the standard for treatment planning and radiologic response measurements in high-grade gliomas,2 whereas our data indicate that the diagnostic performance of CT with contrast to delineate high-grade gliomas before treatment is not necessarily inferior. Clearly, anatomy is better visualized with MR imaging than CT. Furthermore, subtle areas of disease progression that may be outside the main tumor mass are better identified on MR imaging than on CT.
Third, in particular for high-grade gliomas, there is room for improvement in tumor delineation. For instance, radiation oncology guidelines are heterogeneous regarding target delineation.36⇓–38 MR spectroscopy and PET hold promise as additives to the current standard, but availability and standardization are limitations to more widespread use. For a detailed discussion of these techniques, we suggest recent reviews.39⇓⇓⇓–43 Furthermore, T2-weighted imaging performed best for diffuse gliomas as a whole and for low-grade-gliomas but could not be estimated for high-grade gliomas in these data because quantitative information was only available from 2 studies. Nevertheless, T2-weighted imaging may contribute to better delineation of high-grade gliomas as well.
Strengths of this meta-analysis include a thorough search strategy, assessment of reporting quality by STARD criteria, and analysis using the hsROC method, whenever available, with individual sample data.
Our results should be interpreted within the limits of the quality of observational data that were retrieved with the limited number of patients and samples from publications with suboptimal reporting quality. Due to obvious reluctance to sample tissue outside imaging abnormalities, true-negative samples are underrepresented in these data; thus, estimates of false-positive rates are probably less reliable than those of true-positive rates. Furthermore, the available data only allowed indirect comparison of imaging techniques because only 2 studies were identified with quantitative head-to-head comparison of techniques.44,45 Last, several potential biases in this meta-analysis should be considered.46 Methodologic heterogeneity is likely to exist. Publication bias was suggested, though population bias is unlikely because all patients were required to have a glioma, control cases were excluded, and all samples from patients were examined with the same reference standard of histopathologic examination. Nevertheless, verification bias may be present because imaging characteristics have probably guided biopsy sampling strategies. The studies may be biased by patient selection, and we cannot exclude heterogeneity from subjective interpretation of image measurements. Furthermore, clinical heterogeneity is likely because in addition to unstandardized imaging and pathology protocols, positivity criteria of diagnostic and reference tests may have varied among studies.
The implication of our findings is that planning of surgery or radiation therapy for diffuse gliomas using current imaging protocols should be performed with caution because these have only moderate accuracy for glioma delineation based on limited evidence. The sensitivity of imaging to delineate all regions of existing tumor infiltration seems to be less than the specificity to rule out tumor from normal brain. The true-positive rate of conventional imaging for high-grade gliomas may be improved by MR spectroscopy and PET. Furthermore, future effort to quantify and improve this accuracy may aim at combinations of imaging and head-to-head comparison with molecular characterization as the criterion standard.
Conclusions
In this meta-analysis, the diagnostic accuracy of imaging for delineation of diffuse gliomas (low- and high-grade gliomas combined) is best with T2-weighted imaging, followed by MR spectroscopy. The diagnostic accuracy of the common clinically used imaging is better for low-grade gliomas with T2-weighted imaging than for high-grade gliomas with T1-weighted gadolinium-enhanced imaging. Improvement is indicated for high-grade gliomas using advanced imaging techniques, such as MR spectroscopy and PET. Current imaging protocols are based on limited evidence from heterogeneous studies, and future studies with head-to-head comparison and combinations of imaging techniques are required to improve glioma delineation.
Footnotes
Disclosures: Niels Verburg—RELATED: Grant: Dutch Cancer Society resident grant.
This work was supported by a residency fellow grant from the Dutch Cancer Society (nr OAA/H1/VU 2015-7502 to N.V.) and an institutional grant from the Cancer Center Amsterdam (nr 2012-2-05 to P.C.D.W.H.).
Paper previously presented as a poster at: Annual Meeting of the Society for Neuro-Oncology, November 21–14, 2013; San Francisco, California; Annual Meeting of the European Association of Neurosurgical Societies, November 11–14, 2013; Tel Aviv, Israel.
References
- Received January 15, 2017.
- Accepted after revision May 30, 2017.
- © 2017 by American Journal of Neuroradiology