Abstract
BACKGROUND AND PURPOSE: Spiral MR imaging may enable improved image quality and higher scan speeds than Cartesian trajectories. We sought to compare a novel spiral 2D T2-weighted TSE sequence with a conventional Cartesian and an artifact-robust, non-Cartesian sequence named MultiVane for routine clinical brain MR imaging.
MATERIALS AND METHODS: Thirty-one patients were scanned with all 3 sequences (Cartesian, 4 minutes 14 seconds; MultiVane, 2 minutes 49 seconds; spiral, 2 minutes 12 seconds) on a standard clinical 1.5T MR scanner. Three readers described the presence and location of abnormalities and lesions and graded images qualitatively in terms of overall image quality, the presence of motion and pulsation artifacts, gray-white matter differentiation, lesion conspicuity, and subjective preference. Image quality was objectivized by measuring the SNR and the coefficients of variation for CSF, GM, and WM.
RESULTS: Spiral achieved a scan time reduction of 51.9% and 21.9% compared with Cartesian and MultiVane, respectively. The number and location of lesions were identical among all sequences. As for the qualitative analysis, interreader agreement was high (Krippendorff α > .75). Spiral and MultiVane both outperformed the Cartesian sequence in terms of overall image quality, the presence of motion artifacts, and subjective preference (P < .001). In terms of the presence of pulsation artifacts, gray-white matter differentiation, and lesion conspicuity, all 3 sequences performed similarly well (P > .15). Spiral and MultiVane outperformed the Cartesian sequence in coefficient of variation WM and SNR (P < .01).
CONCLUSIONS: Spiral 2D T2WI TSE is feasible for routine structural brain MR imaging and offers high-quality, artifact-robust brain imaging in short scan times.
ABBREVIATIONS:
- CV
- coefficient of variation
- SENSE
- sensitivity encoding
T2WI sequences are essential components of every clinical MR imaging protocol. Specifically, the T2 contrast is particularly useful to analyze brain anatomy, CSF spaces, and parenchymal lesions.
Currently, most clinical institutions rely on axially acquired cartesian 2D T2WI TSE sequences.1 These sequences offer a relatively short scan time (especially when accelerated by means of parallel imaging2 or compressed sensing3) and achieve a reliable and accurate depiction of the brain. Alternatively, especially in pediatric and elderly patients, certain institutions rely on sequences with a non-Cartesian sampling scheme, such as PROPELLER.4,5 With PROPELLER, multiple echo-trains of TSE are acquired in a rotating, partially overlapping fashion rather than in a rectilinear fashion as in Cartesian imaging.5 While sequences with this sampling scheme offer central k-space oversampling and thus increased robustness toward artifacts, they require a considerable increase in scan time compared with their Cartesian counterparts in case of fully matching scan parameters.5 Thus, a sequence offering both artifact robustness and an intrinsically short scan time is highly desirable for clinical T2-weighted brain MR imaging.
Despite requiring high technological standards both in terms of scanner hardware and software, spiral MR imaging sequences have recently been implemented for clinical MR imaging. Sequences with spiral k-space sampling have inherently reduced gradient moments, central k-space oversampling, and a nondedicated phase-encoding direction, thus rendering them less susceptible to artifacts. Furthermore, the efficiency of k-space sampling is high in spiral MR imaging due to the longer acquisition duration per shot.6 Recent clinical feasibility studies have thus shown the value of spiral MR imaging for T1-weighted spin-echo/gradient recalled-echo brain and spine imaging as well as for intracranial vessel imaging.6⇓⇓⇓⇓⇓⇓-13
Here we expand on previous endeavors by presenting, for the first time, initial clinical results of a spiral TSE technique for routine clinical 2D T2-weighted TSE brain MR imaging. We hypothesized that this sequence would offer both short scan times and artifact robustness, thus combining the best features of conventional Cartesian imaging and non-Cartesian imaging. To this extent, we prospectively compared the novel spiral 2D T2WI TSE sequence with a conventional Cartesian 2D T2WI TSE sequence and a (non-Cartesian) PROPELLER-like sampled 2D T2WI TSE sequence in patients.
MATERIALS AND METHODS
Study Design and Subjects
This institutional review board-approved intraindividual comparison study was performed between January and May 2021. All participants gave general written informed consent. We prospectively acquired all 3 sequences in 31 consecutive patients (mean age, 57 years; age range, 18–85 years; 16 men, 15 woman) who were referred to our department for brain MR imaging due to various clinical indications. Exclusion criteria were as follows: general contraindication to MR imaging (ie, metallic implants and so forth), younger than 18 years of age, and pregnancy. The final main diagnoses were the following: chronic lacunar infarcts (n = 1), periventricular heterotopia (n = 1), territorial infarcts (n = 6), metastases (n = 1), microangiopathy (n = 9), multiple sclerosis demyelinating lesions (n = 1), periventricular leukomalacia (n = 1), subdural hematoma (n = 1), dilated Virchow-Robin spaces (n = 7), and normal findings (n = 3).
MR Imaging
Imaging was performed on a standard clinical 1.5T scanner (Ingenia; Philips Healthcare) with a 16-channel head coil. The combination of spiral with TSE was enabled with the Compressed SENSE 2.0 WIP software, Release 5.7 (Philips Healthcare). As part of the routine clinical protocol, the following additional sequences were acquired besides the 2D T2WI TSE scans: a sagittal 3D FLAIR sequence, a precontrast sagittal 3D T1WI turbo field echo sequence, a diffusion-weighted transverse sequence, a susceptibility-weighted transverse sequence, and, in selected cases, a sagittal postcontrast 3D T1 black-blood TSE sequence or a postcontrast sagittal 3D T1 m-Dixon turbo field echo sequence.
The 3 2D T2WI TSE sequences (Table 1) were acquired in random order. The performance of the spiral sequence was benchmarked against 2 commercially available sequences used routinely at our institution: A conventional Cartesian 2D T2WI TSE sequence and a MultiVane 2D T2WI TSE sequence using PROPELLER-like, non-Cartesian k-space sampling. Sequence parameters of the Cartesian and MultiVane sequences were chosen on the basis of long-standing and well-established clinical protocols.10 The sequence parameters of the spiral sequence were selected on the basis of the initial optimization of this sequence by the vendor and in-house by means of volunteer tests. Most important, the sequence parameters were kept as constant as possible among all 3 sequences.9⇓-11 However, due to institutional scan time constraints, MultiVane was accelerated with sensitivity encoding (SENSE), and the number of signal averages was adapted to shorten the scan time.
Sequence parameters
The spiral sequence uses an in-plane spiral in-out readout scheme (spiral acquisition window, 11.4 ms; scan duration, 2 minutes 12 seconds). Blurring due to off-resonance was corrected during reconstruction on the basis of a magnetic field map acquired before the spiral scans.11 This study was performed on the standard product configuration without additional enhancement. Eddy current calibration of B0 eddy currents and linear and cross-term eddy currents was performed as part of the standard system tuning procedure. Compensation of those eddy current contributions was performed in run-time as part of the vendor's product-acquisition software.14
Qualitative Analysis
Image analysis was performed independently by 3 readers (a board-certified neuroradiologist with 30 years of experience and 2 trainees, each with 3 years of experience in medical imaging) in a blinded and randomized manner.
First, as outlined elsewhere,15 for each sequence and patient, the readers recorded the presence, number, and localization of abnormalities and lesions. In case of discrepancies among the 3 sequences or among the readers, the lesions were evaluated on all available imaging sequences in consensus to detect potential false-positive and false-negative findings.15
Second, as suggested elsewhere,16⇓⇓-19 all images were graded one-by-one in the following categories using 4-point Likert scales: overall image quality (1, nondiagnostic; 2, limited but interpretable; 3, minimally limited; and 4, optimal quality), the presence of motion and pulsation artifacts (1, severe image artifacts; 2, moderate artifacts; 3, mild artifacts; 4, no artifacts), gray-white matter differentiation (1, indistinguishable gray-white sharpness; 2, very blurry gray-white sharpness; 3, slightly blurry gray-white sharpness; 4, well-defined gray-white sharpness), and lesion conspicuity (1, a lesion whose borders are barely distinguishable from background brain; 2, a lesion with very blurry margins; 3, a lesion with slightly blurry margins; 4, sharp lesion margins). Additionally, readers were asked to record any other artifacts or anything unusual that appeared to them during the readout.
Last, for each patient, all 3 sequences were presented side-by-side, and the readers were asked to order the sequences according to their subjective preference. Specifically, a score of 3 was given to the best sequence; a score of 2, to the second-best sequence; and a score of 1, for the worst sequence. Scores could be used more than once if sequences were judged equivalent.20
Quantitative Analysis
As secondary end points, we used ROI-based analyses to objectify image quality and image appearance.9,16 For each patient, ROIs were drawn on representative images from each sequence. ROIs were positioned within the frontal WM, the GM of the caudate head, and in the lateral ventricles (CSF) at the level of foramen of Monro.17 All ROIs were the same size and had identical positioning between sequences.16 From each ROI, the mean signal intensity and SD of the signal intensity were extracted. ROI placement was performed twice, and the average values of both measurements were considered representative for further analysis.9 As suggested in the literature,9,17,21 we then computed the SNR and the coefficient of variations (CVs) for WM, GM, and CSF as follows:
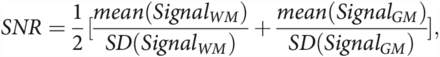
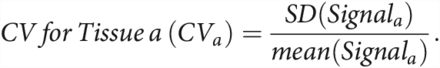
Statistical Analysis
Data distribution was initially checked with histograms, boxplots, and quantile-quantile plots. Differences in qualitative metrics were initially checked with Friedman tests followed by post hoc Wilcoxon signed rank tests. Interreader agreement of qualitative scores was quantified with Krippendorff α coefficients (0, no agreement, 1, perfect agreement). Differences in quantitative metrics were initially checked with 1-way repeated-measures ANOVAs followed by post hoc paired t tests. P values were corrected for multiple comparisons with the Holm method. P values < .05 were considered significant. All statistical analyses were performed in the R programming language (http://www.r-project.org/).
RESULTS
Representative image examples are shown in the Figure and the Online Supplemental Material. With a scan time of 02:12 minutes, spiral achieved a scan time reduction of 51.9% and 21.9% compared with Cartesian and MultiVane, respectively.
Representative images from 3 different patients (1 patient per row). In patient 1, ghosting artifacts from bulk motion appear on Cartesian images (orange arrows) but not on MultiVane or spiral images. In patient 2, no artifacts were present and thus an excellent visualization could be achieved for all 3 sequences. In patient 3, a small focal multiple sclerosis lesion is visualized clearly on all 3 sequences (green arrow).
In terms of the presence, number, and location of lesions, there were no differences among the 3 sequences. All readers recorded the same pathologic and abnormal findings on all 3 sequences. Besides motion and pulsation artifacts, no other types of artifacts were recorded.
Concerning the qualitative metrics (Table 2 and Online Supplemental Material), interreader agreement was high (α = .755 for presence of pulsation artifacts; α = .98 for presence of motion artifacts). Spiral and MultiVane both outperformed the Cartesian sequence in terms of overall image quality and the presence of motion artifacts (P < .001 for all readers and both metrics). Between spiral and MultiVane, there were no significant differences in these 2 metrics (P > .4 for all readers and both metrics). In terms of the presence of pulsation artifacts and gray-white matter differentiation, all 3 sequences performed similarly well (P > .15 for each reader). Finally, for lesion conspicuity, there were also no significant differences among the 3 sequences (P > .45 for each reader). Readers indicated a higher subjective preference for spiral and MultiVane compared with the Cartesian sequence (P < .001 for all readers) without significant differences between spiral and MultiVane (P > .5 for all readers).
Detailed overview of qualitative and quantitative data
Concerning the quantitative metrics (Table 2 and Online Supplemental Material), there were no significant differences among the 3 sequences for the metrics CVCSF and CVGM (P > .12 for both metrics). However, both spiral and MultiVane outperformed the Cartesian sequence in the metrics CVWM and SNR (P < .01 for both sequences and metrics). Between spiral and MultiVane, there were, however, no differences in terms of CVWM and SNR (P > .5 for both metrics).
DISCUSSION
In this study, we compared a novel spiral 2D T2WI TSE sequence with its conventional Cartesian counterpart and an artifact-robust, PROPELLER-like sampled sequence named MultiVane. We showed that the spiral sequence outperforms the Cartesian sequence and performs equally as well as the MultiVane sequence in terms of subjective and objective image quality. Concerning the exact frequency and nature of artifacts, all 3 sequences exhibited only motion and pulsation artifacts. However, the spiral and MultiVane both had significantly fewer motion artifacts than the Cartesian sequence, while all 3 sequences had only minor pulsation artifacts in select cases. Thus, the spiral sequence enables high-quality, artifact-robust imaging on a level with the MultiVane sequence, yet at a shorter scan time.
Spiral MR imaging has several benefits over Cartesian k-space sampling. Due to the longer acquisition duration per shot, scan efficiency is high and the scan time can, thus, be very short. Furthermore, spiral trajectories show inherently reduced gradient moments, central k-space oversampling, and a nondedicated phase-encoding direction. These traits render spiral sequences more robust toward artifacts. Thus, with spiral MR imaging, rapid and artifact-robust imaging can be achieved.6 Accordingly, promising clinical results have been reported for anatomic spiral T1WI spin-echo and gradient recalled-echo imaging. While the technical details of spiral TSE and T2WI imaging have been described previously,22,23 to the best of our knowledge, this is the first study investigating the value of a spiral TSE technique as implemented for 2D T2-weighted TSE brain MR imaging in patients on a standard clinical MR imaging scanner and clinical routine.
Currently, most institutions rely on Cartesian or PROPELLER-like sampled sequences (such as MultiVane) for clinical T2-weighted brain MR imaging. While PROPELLER sequences have become a popular choice for anatomic brain imaging due to their increased robustness predominantly toward motion artifacts, scan time is generally increased compared with conventional Cartesian TSE imaging.5,24,25
A standard rectilinear sequence requires (M/L) excitations to fill a k-space of matrix size M with acquisitions of echo-train length L. For PROPELLER imaging, at least 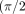 ) × (M/L) excitations are needed for an equivalent sequence, thus leading to approximately 60% increase in scan time compared with Cartesian imaging.5 With parallel imaging or compressed sensing acceleration, the increase in scan time can be reduced. Specifically, as in the clinical routine at our institution, our MultiVane sequence also used SENSE 1.5 and 1 signal average, explaining why MultiVane had a shorter scan time than the Cartesian sequence. Hypothetically, if the MultiVane sequence had been acquired without SENSE and with 2 signal averages, as in the case of the Cartesian and spiral sequences, scan time would have increased by a factor of 2.5, which would have resulted in a 66% increase in scan time compared with the Cartesian sequence.
) × (M/L) excitations are needed for an equivalent sequence, thus leading to approximately 60% increase in scan time compared with Cartesian imaging.5 With parallel imaging or compressed sensing acceleration, the increase in scan time can be reduced. Specifically, as in the clinical routine at our institution, our MultiVane sequence also used SENSE 1.5 and 1 signal average, explaining why MultiVane had a shorter scan time than the Cartesian sequence. Hypothetically, if the MultiVane sequence had been acquired without SENSE and with 2 signal averages, as in the case of the Cartesian and spiral sequences, scan time would have increased by a factor of 2.5, which would have resulted in a 66% increase in scan time compared with the Cartesian sequence.
With spiral MR imaging, however, the dilemma of artifact robustness and scan speed can be resolved. Specifically, our spiral sequence offers exceptionally short scan times as well as both motion and pulsation artifact robustness, thus combining the best features of conventional Cartesian imaging and non-Cartesian (PROPELLER-like) imaging. In this context, the spiral sequence achieved a scan time reduction of nearly 50% compared with the Cartesian sequence. Most important, while not investigated in this study, scan time for the spiral sequence may be shortened even further with parallel imaging or compressed sensing techniques.
A further important aspect of the current study concerns the exact choice and configuration of the sequences. While 2D TSE techniques are widely considered the reference standard for clinical brain MRI,1 3D and multiband TSE techniques have also been proposed for clinical T2-weighted brain imaging. While not investigated in this study, our spiral TSE technique is highly adaptable and may be combined with these approaches.22,26,27 Furthermore, with high-field imaging being popular for brain MR imaging, our spiral sequence can also be acquired at 3T. Thus, our spiral sequence had lower specific absorption rate values than its counterparts, an advantage at 3T. However, the spiral sequence is more susceptible to off-resonance B0 effects, which are more pronounced at 3T. Thus, a future study assessing the clinical value of our spiral TSE technique at 3T would be of interest.
One limitation of our study was the limited sample size: Because the spiral sequence could not be run with standard clinical software, the scanner console had to be rebooted before each spiral acquisition (to load a patch). This step limited our ability to acquire the sequence in further patients. Second, it may have not been possible to fully blind readers toward sequence details because the spiral sequence has a distinct appearance. Third, scan parameters were not fully identical among all sequences, possibly representing a source of bias, especially for the quantitative image analysis. Last, we are aware that while formula-based approaches for estimation of quantitative metrics are widely used, they are inherently limited, and more sophisticated methods may yield more accurate estimations.9
CONCLUSIONS
We show that a spiral 2D T2WI TSE sequence enables high-quality, artifact-robust brain MR imaging in clinical routine at short scan times. This sequence may, thus, represent a promising option for improved and rapid clinical T2-weighted brain MR imaging.
Footnotes
Disclosure forms provided by the authors are available with the full text and PDF of this article at www.ajnr.org.
References
- Received June 5, 2021.
- Accepted after revision July 23, 2021.
- © 2021 by American Journal of Neuroradiology